Ambient DNA Preservation Without Compromising Sequencing: Our Role in Evaluating Ensilication
Whole‑genome analysis of storage-induced artifacts in tumor and normal DNA
At Fulcrum Genomics, we care a lot about what happens before variant calling: how samples are collected, stored, and prepared. A single artifactual C>T mutation in the wrong place can look identical to a cancer driver mutation. If sample storage and preservation introduces thousands of these artifacts, no amount of downstream pipeline tuning will fix it.
“Identifying and eliminating sources of error has been central to Fulcrum’s work since our inception,” said Nils Homer, Founding Partner at Fulcrum Genomics. “This work shows that ensilication can preserve samples at room temperature while actually reducing the artifacts that complicate variant interpretation.”
Why DNA Storage Still Matters
Today, most nucleic acid preservation depends on cold storage, typically from −20 °C to −80 °C, and sometimes even colder. That infrastructure is expensive, energy-intensive, and hard to scale globally. As precision oncology and population genomics expand, we need preservation methods that don’t require a dense network of ultra‑low temp freezers and the associated cold chain.
As Michael Becich, CEO of CacheDNA, puts it: “The first era of genomics was built on cold storage. The next belongs to elegant biochemistries that preserve truth at the molecular level without ongoing energy or intervention. Every sample tells a story, and we believe that if a sample can be collected, it should be carefully protected until ready for testing. We’re building the bridge that makes modern biopreservation possible to reveal those insights.”
Ambient-temperature solutions like Cache DNA’s ensilication are promising, but they must clear a high bar:
Maintain clinical variant detection
Avoid introducing artifactual mutations
Preserve DNA integrity across temperatures and time
This study tackles those questions directly using clinical FFPE tumor samples, targeted mutation assays, and deep whole‑genome sequencing.
What the Study Shows
The study evaluates ensilication side-by-side with standard −80 °C storage across multiple dimensions:
Clinical panels showed perfect concordance.
Lung cancer DNA extracted from FFPE blocks was split, stored either frozen or ensilicated at room temperature for 14 days, and then tested using a routine clinical panel. Across ten samples, there was 100% diagnostic concordance between storage methods, including low VAF variants down to ~2%, meaning no gained or lost actionable mutations after 14 days of storage.
Whole‑genome sequencing revealed frozen storage introduced more artifacts.
For 32 matched tumor–normal pairs, DNA aliquots were stored frozen (−80 °C) or ensilicated at room temperature for 28 days and then sequenced (30x normal, 80x tumor). Overall sequencing metrics, including insert sizes, chimera rates, duplication rates, and library complexity, were comparable between storage conditions, enabling direct comparison of storage-induced mutations.
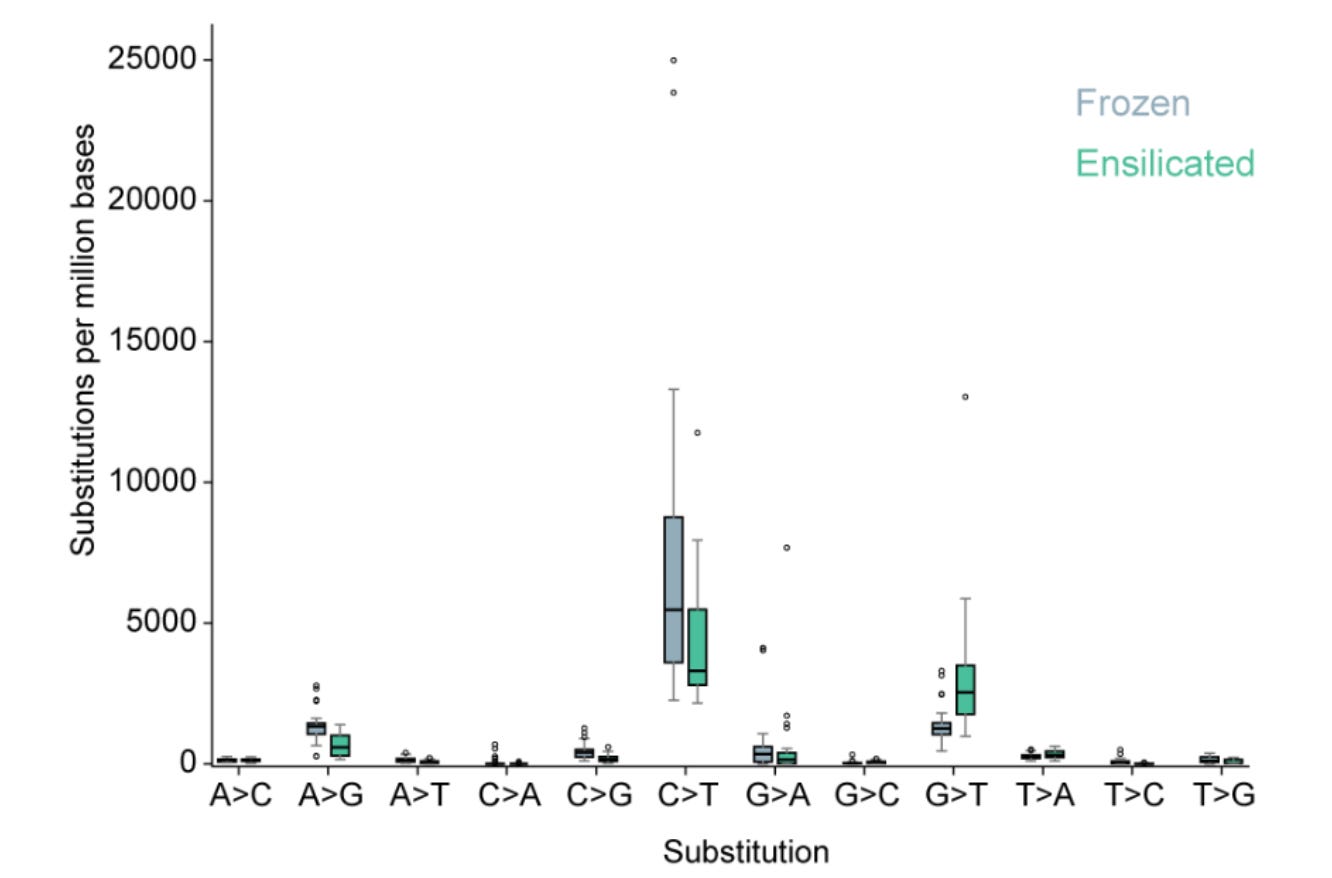
Frozen −80 °C storage of FFPE tumor DNA generated substantially more artifactual C>T mutations than ensilication at room temperature, increasing background ‘noise’ that can mimic true cancer variants.
The key finding: frozen storage accumulated substantially more artifactual C>T mutations in tumor DNA than ensilication, particularly in CpG contexts. Frozen samples showed up to 65% more C>T substitutions per million bases than ensilicated samples, adding thousands of apparent mutations that are indistinguishable from true somatic events. These artifacts are especially concerning at clinically relevant CpG sites frequently mutated in cancer.
Ensilication worked across DNA shapes and extreme temperature ranges.
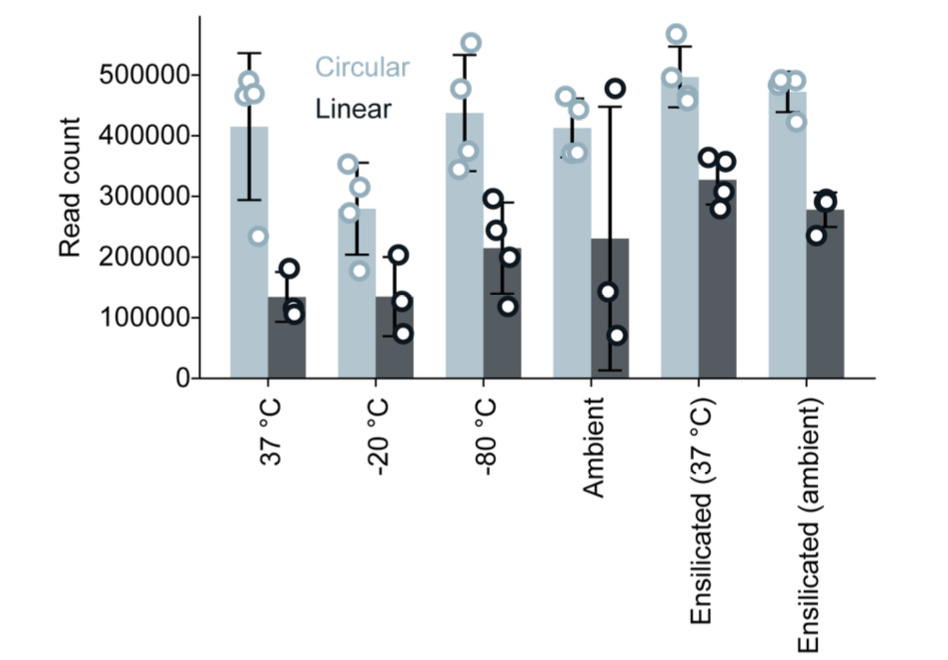
The study examined both linear ssDNA and circular ssDNA (cssDNA) libraries across a range of temperatures (−80 °C to 37 °C) with and without ensilication over 20 days. Circular libraries showed strong inherent stability even unprotected, while linear DNA benefited markedly from encapsulation, with ensilicated linear samples outperforming unprotected controls across every tested temperature.
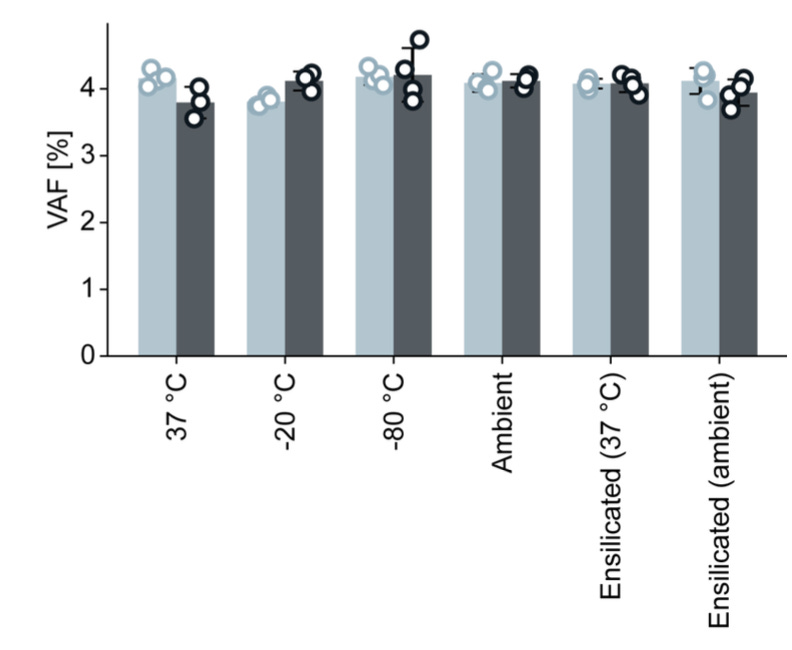
Across −80 °C to 37 °C, circular ssDNA libraries remained highly stable, and ensilication markedly improved the stability of linear ssDNA, while all conditions still supported accurate detection of ~5% variants.
Importantly, even where some loss of amplifiable molecules occurred, variants above 5% VAF remained accurately detectable in all conditions and topologies.
Fulcrum Genomics’ Contributions
Fulcrum’s role in this work was to make the storage comparison quantitative at whole‑genome scale, not just qualitative at a few loci. This included:
Processing and aligning whole‑genome data from matched tumor–normal pairs, ensuring that differences between storage methods weren’t confounded by pipeline noise.
Calling and filtering somatic variants to separate true tumor mutations from storage-induced artifacts that could affect downstream interpretation.
Analyzing and visualizing substitution rates and sequence context to reveal the characteristic patterns of damage associated with frozen vs. ensilicated storage.
By standardizing these analyses across all samples, we were able to show not just that ensilication preserves clinical calls, but also how different storage conditions reshape the apparent mutation landscape, especially the excess C>T artifacts seen in frozen FFPE tumor. That level of detail is critical for applications like tumor mutational burden, signature analysis, and regulatory decision‑making.
“Our goal has always been maximum sensitivity without sacrificing accuracy for clinical decision-making. This work with Cache DNA represents a significant step in that direction, showing Cache DNA’s ensilication technology can preserve samples at room temperature while actually reducing the artifacts that complicate variant interpretation,” said Homer.
This kind of artifact‑aware, whole‑genome benchmarking is a natural extension of our work: helping teams understand not only what their sequencing data says, but how upstream choices—from preservation through library prep—shape the answers they get.
Working with James Banal, Co-Founder of Cache DNA, and the broader team was a model collaboration. Their rigorous experimental design made the computation analysis straightforward to interpret and the findings unambiguous.
Implications for Genomic Medicine
The findings suggest that ambient ensilication can match or outperform −80 °C storage in preserving sequencing fidelity, at least over the timeframes evaluated. That matters for:
Clinical oncology – Reducing storage-induced artifacts lowers the risk of false-positive somatic calls, especially in FFPE samples where C>T damage is already endemic. This study directly evaluated clinically relevant genomic biomarkers to ensure downstream clinical decisions rest on robust data.
Regulatory and biomarker development – Artifact-aware preservation supports reliable measurement of tumor mutational burden, mutational signatures, and subclonal architecture. Removing temperature requirements also opens opportunities for global drug development programs that rely on sample transit to centralized labs.
Global and decentralized genomics – Ambient storage reduces cost and infrastructure barriers, making it feasible to collect and ship samples from remote or resource-limited settings without sacrificing data quality.
Of course, more work is needed: multi‑year studies, diverse tissue types, and broader operational and economic analyses will be important to understand where ambient methods can replace or complement cold storage at scale. But this study provides strong, whole‑genome evidence that ensilication is a viable path toward cold‑chain‑free genomic medicine.
At Fulcrum Genomics, we’re glad to partner with teams like Cache DNA on this kind of foundational work where careful computational analysis helps clarify not just what variants we see, but why we see them, and whether they’re real.
As ensilication and other ambient preservation methods mature, we’re committed to bringing the same rigor to evaluating them that we’ve applied to library preparation, sequencing platforms, and variant calling. Because genomic medicine is only as good as the samples it starts with.
📖Read the paper:
Evaluation of ensilication technology for ambient DNA preservation
💻Code and data:
Analysis code is available at Zenodo (10.5281/zenodo.17469053); sequencing data will be deposited to EGA upon publication.